Building a Human Genotype-Phenotype Map
Genome-wide association studies have mapped thousands of genetic associations, but interpreting what those associations mean biologically remains a central challenge. In a new medRxiv preprint, Andrew Elmore, Aimee Hanson, Genevieve Leyden and colleagues introduce the Human Genotype-Phenotype Map (GPMap), an open resource for tracing shared genetic signals across complex traits and molecular measurements.
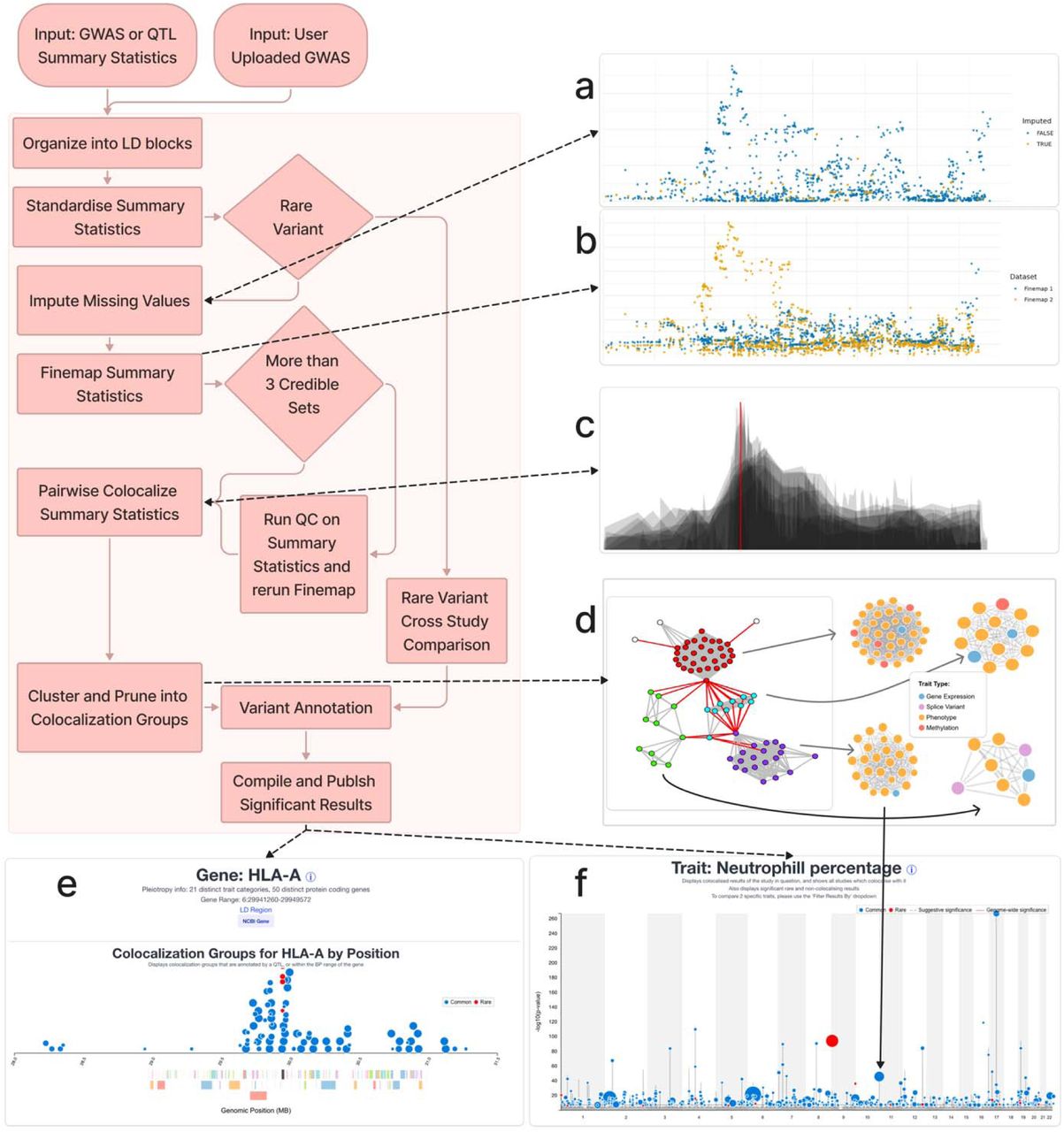
Figure: GPMap processing pipeline, from GWAS summary statistics through imputation, fine-mapping, colocalisation and views by trait, gene, variant and tissue. Source: Elmore et al., medRxiv, 2026, Fig. 1 (CC BY-ND 4.0).