Building a Human Genotype-Phenotype Map
Genome-wide association studies have mapped thousands of genetic associations, but interpreting what those associations mean biologically remains a central challenge. In a new medRxiv preprint, Andrew Elmore, Aimee Hanson, Genevieve Leyden and colleagues introduce the Human Genotype-Phenotype Map (GPMap), an open resource for tracing shared genetic signals across complex traits and molecular measurements.
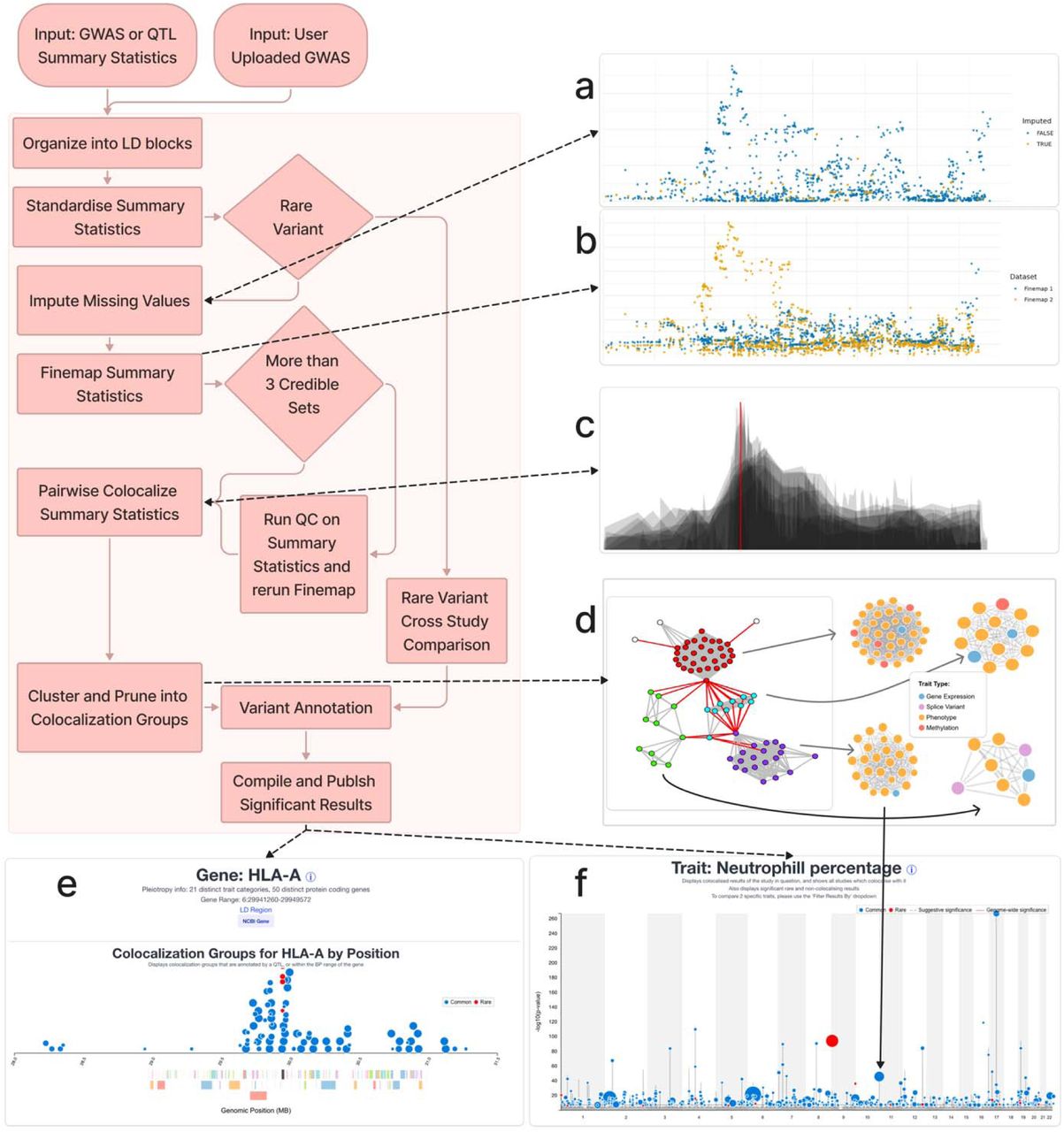
Figure: GPMap processing pipeline, from GWAS summary statistics through imputation, fine-mapping, colocalisation and views by trait, gene, variant and tissue. Source: Elmore et al., medRxiv, 2026, Fig. 1 (CC BY-ND 4.0).
Background
A single genetic variant can associate with multiple traits. Sometimes this reflects a causal chain; sometimes it reflects a shared molecular mechanism; sometimes it reflects distinct nearby signals that are difficult to separate. Colocalisation helps distinguish these possibilities by asking whether two associations are likely to share the same causal variant.
Utilising the extensive OpenGWAS data resource, GPMap scales this idea across a very large space of traits and molecular measurements, aiming to make pleiotropic structure easier to search, cluster and use.
What the study did
The preprint describes a repository of colocalising genetic associations across 15,997 complex traits and 2.7 million molecular measurements. The resource integrates common and rare variant evidence, cis- and trans-acting effects, tissue-specific measurements and single-cell datasets.
The processing pipeline includes summary statistic imputation, fine-mapping, colocalisation and graph-based clustering of shared signals. The resulting data can be explored by trait, gene, variant or tissue through the GPMap website, an Application Programming Interface (API), R package and downloads.
What this adds
The authors report more than 49.3 million colocalising trait pairs, aggregated into 97,393 colocalisation groups. They also report that 55.8% of genome-wide significant disease-associated loci colocalise with at least one molecular trait.
The preprint illustrates how this structure can help refine complex phenotypes, including tissue-informed dissection of BMI-associated loci. It also shows how colocalisation evidence can support drug target prioritisation and improve the selection of genetic instruments for causal inference.
Why this matters
The GPMap is designed as a map of potential hypothesis space rather than addressing a single hypothesis test. Its value is in making shared genetic architecture searchable at scale, so that researchers can move from a GWAS signal to plausible molecular context much more rapidly.
However, this does not remove the need for careful interpretation. Colocalisation is evidence of a shared genetic signal, not automatic proof of a biological mechanism or clinical utility. But as the number of GWAS and molecular QTL datasets keeps growing, systematic resources like GPMap can make genetic evidence more usable for epidemiology, functional genomics and drug target prioritisation.
This work is currently a preprint and has not yet been peer reviewed.
Preprint and resource
Elmore AR, Hanson AL, Leyden GM, Johnson J, Davey Smith G, Paternoster L, Gaunt TR, Hemani G. Building The Human Genotype-Phenotype Map to Harness Pleiotropy and Refine Disease Mechanisms. medRxiv (posted 2026-02-20). DOI: 10.64898/2026.02.19.26346618.
GPMap resource: https://gpmap.opengwas.io