Using genetics to prioritise therapeutic targets for immune-mediated diseases
Immune-mediated diseases such as asthma, eczema, inflammatory bowel disease, rheumatoid arthritis and multiple sclerosis share parts of the same immune biology, but translating that biology into therapeutic targets is still difficult. In a new paper in Scientific Reports, Maria Sobczyk and Tom Gaunt use integrative Mendelian randomization (MR) approaches to evaluate potential drug targets across 14 immune-mediated diseases.
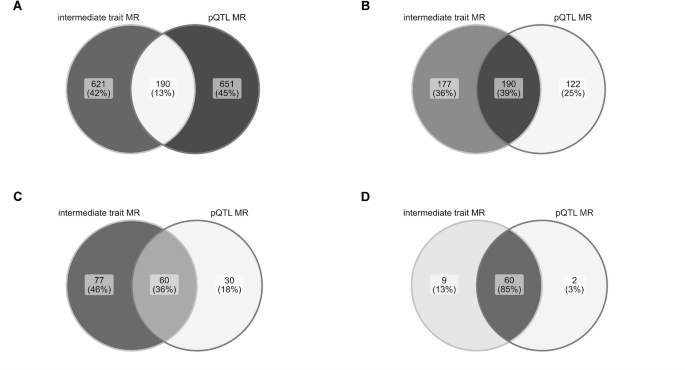
Figure: Overlap between immune-cell-informed MR and protein-QTL MR evidence for gene-immune-mediated disease associations, illustrating why combining molecular layers adds information beyond either approach alone. Source: Sobczyk and Gaunt, Scientific Reports, 2026, Fig. 4 (CC BY 4.0).
Background
Human genetic evidence can help prioritise drug targets because inherited variation provides a natural perturbation of biological pathways. For immune-mediated diseases, this is especially useful: immune cell abundance, circulating proteins and disease risk can all be connected using large-scale genome-wide association studies.
The challenge is that different molecular measurements capture different aspects of biology. A target supported by immune-cell genetics may not be the same as one supported by circulating protein abundance. This study asks how those evidence sources compare, and what can be learned by combining them.
What the study did
The analysis focused on 834 targets that have been investigated for immune-mediated disease treatment. The authors used two complementary MR strategies:
- immune-cell-informed MR, using genetic instruments related to peripheral blood immune cell abundance;
- protein-QTL MR, using genetic instruments for circulating protein abundance from large plasma proteomic studies.
Both sets of MR results were followed by colocalisation analyses, helping to distinguish signals that are more likely to reflect a shared causal variant from those that may arise because nearby variants are correlated through linkage disequilibrium.
What this adds
The study first identified genetic evidence for relationships between major immune cell types and immune-mediated diseases, including strong evidence involving eosinophils in atopic and inflammatory conditions.
It then used these immune-cell and protein-QTL instruments to prioritise gene-disease and protein-disease associations. The two approaches produced partially overlapping, but substantially complementary, sets of candidate targets. That limited overlap is itself informative: relying on one molecular layer alone would miss a sizeable part of the target-prioritisation landscape.
The paper also compares genetic evidence with approved and in-development drug indications, highlighting examples where the genetic evidence supports known biology and where it suggests possible repurposing opportunities. These results should be treated as prioritisation evidence rather than direct proof of clinical efficacy.
Why this matters
Drug repurposing in immune-mediated disease is attractive because many targets and pathways are shared across conditions. MR and colocalisation cannot replace experimental pharmacology or clinical trials, but they can help decide which target-disease pairs deserve more attention.
The main practical message is that target prioritisation benefits from triangulation. Immune-cell traits and protein abundance are not interchangeable measurements, and combining them gives a broader view of plausible therapeutic biology.
Paper
Sobczyk MK, Gaunt TR. Integrative mendelian randomization approaches for therapeutic target prioritisation in immune-mediated diseases. Scientific Reports (2026) 16:11851. DOI: 10.1038/s41598-026-41818-3. PubMed: 41775781. Full text: PMC13065838.