Using genetics to prioritise therapeutic targets for immune-mediated diseases
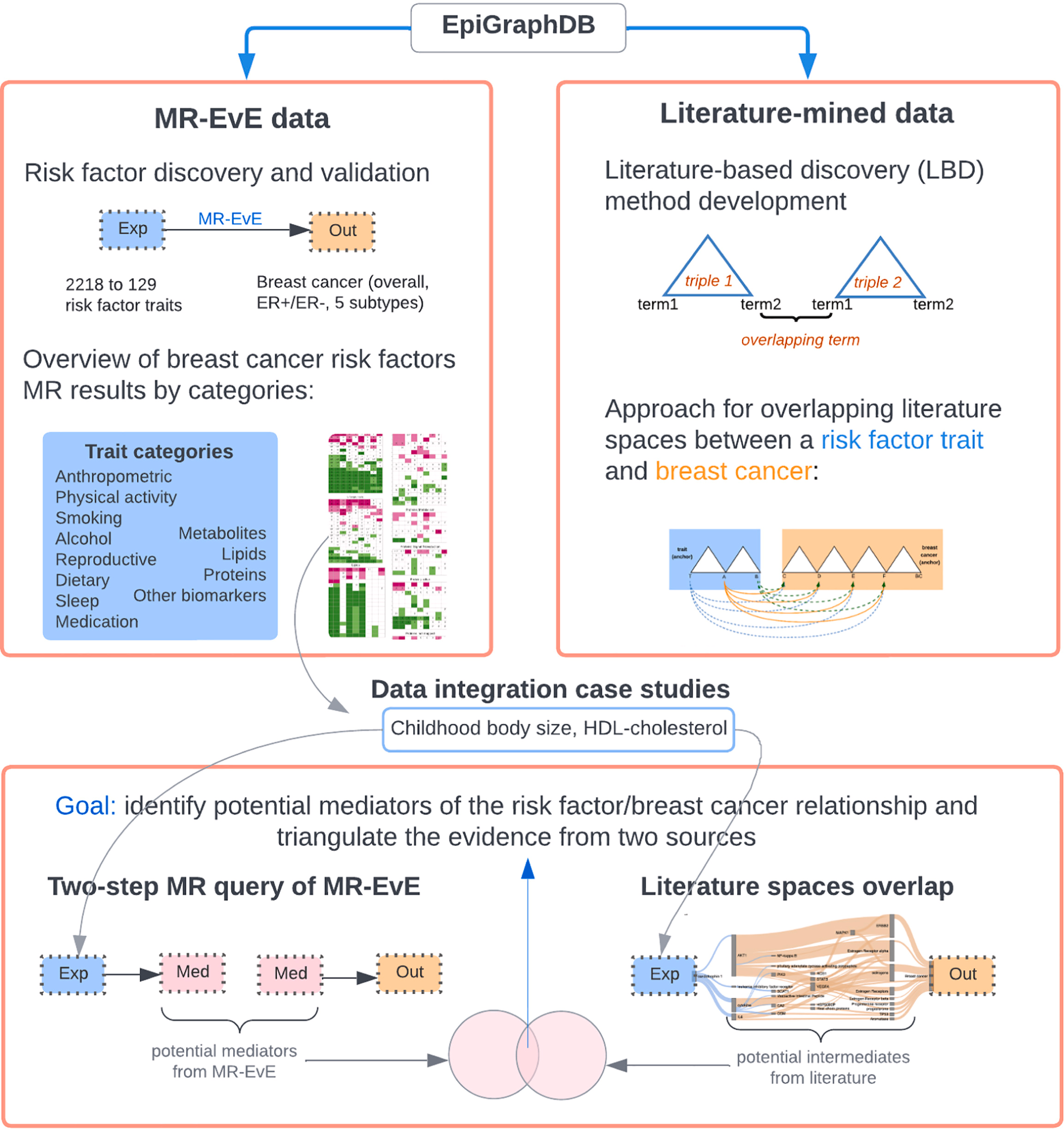
Immune-mediated diseases such as asthma, eczema, inflammatory bowel disease, rheumatoid arthritis and multiple sclerosis share parts of the same immune biology, but translating that biology into therapeutic targets is still difficult. In a new paper in Scientific Reports, Maria Sobczyk and Tom Gaunt use integrative Mendelian randomization (MR) approaches to evaluate potential drug targets across 14 immune-mediated diseases.
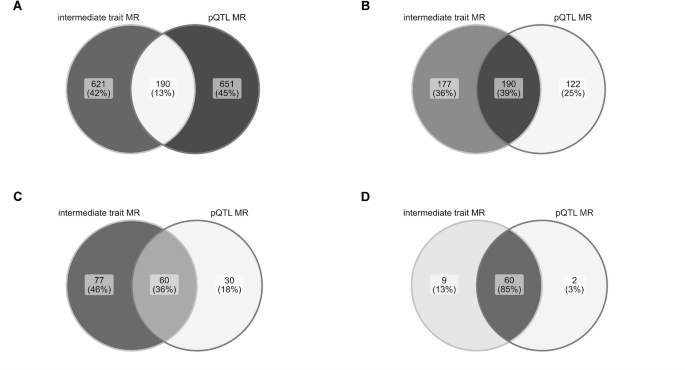
Figure: Overlap between immune-cell-informed MR and protein-QTL MR evidence for gene-immune-mediated disease associations, illustrating why combining molecular layers adds information beyond either approach alone. Source: Sobczyk and Gaunt, Scientific Reports, 2026, Fig. 4 (CC BY 4.0).