Mapping age-dependent genetic influences on DNA methylation
DNA methylation is often analysed as though genetic effects are stable over time, but development and ageing may change how genetic variation influences the methylome. In a new bioRxiv preprint, Yueying Li and colleagues map age-dependent methylation quantitative trait loci (mQTLs) using repeated DNA methylation measures from birth through adulthood.
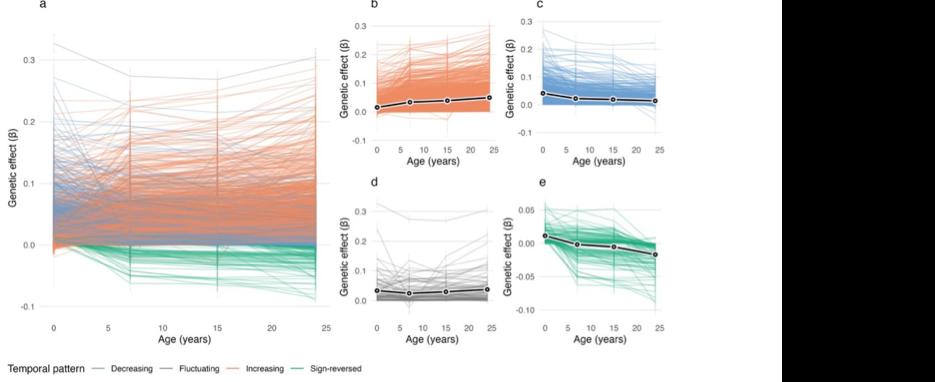
Figure: Trajectories of longitudinal mQTL-CpG associations, grouped by temporal pattern, showing genetic effects that increase, decrease, fluctuate or change sign across age. Source: Li et al., bioRxiv, 2026, Fig. 3 (CC BY 4.0).
Background
mQTL studies connect genetic variants to DNA methylation levels at nearby or distant CpG sites. These analyses are useful for understanding gene regulation, interpreting epigenome-wide association studies and connecting genetic variation to molecular mechanisms.
Most mQTL studies use cross-sectional data. That makes it hard to distinguish stable genetic effects from effects that become stronger, weaker or otherwise change across the life course. Longitudinal methylation data provide a way to ask that question directly.
What the study did
The authors used repeated blood DNA methylation measures from the Avon Longitudinal Study of Parents and Children, spanning birth, childhood, adolescence and adulthood. They applied linear mixed models to identify genotype-by-age effects on DNA methylation.
They then compared the resulting longitudinal mQTL effects with evidence from two independent cohorts: the Generation R Stud and the Drakenstein Child Health Study. This was important because age-dependent genetic effects could be sensitive to cohort structure, ancestry, cell composition and measurement context.
What this adds
The preprint reports 2,210 longitudinal mQTLs, corresponding to 2,393 SNP-CpG pairs. About half of these loci showed increasing genetic effects with age. The reported genotype-by-age effects were directionally consistent in the two independent cohorts, with stronger agreement in Generation R and more modest but still positive agreement in Drakenstein.
CpGs with longitudinal mQTLs were more heritable and enriched in regulatory elements and pathways related to development and cell adhesion. The results suggest that genetic regulation of DNA methylation is not always a fixed property measured once and carried forward unchanged.
Why this matters
For life-course epidemiology and molecular genetics, this is a useful reminder that timing matters. A genetic variant may have a weak effect on methylation at one age and a stronger effect later, or vice versa. That has implications for interpreting epigenetic associations, building methylation-based biomarkers and selecting molecular instruments for downstream causal analyses.
The work is currently a preprint and has not yet been peer reviewed. Its central contribution is a resource and framework for studying dynamic genetic regulation of DNA methylation, rather than a claim that individual longitudinal mQTLs are immediately clinically actionable.
Preprint
Li Y, Staley JR, Grossbach A, Cappadona C, Lussier AA, Dunn EC, Felix JF, Cecil CAM, Stein DJ, Zar HJ, Walker V, Gaunt TR, Tilling K, Mulder R, Simpkin AJ, Hemani G, Min JL. Identification of Dynamic Genetic Influences on DNA Methylation from Birth to Adulthood. bioRxiv (posted 2026-03-10). DOI: 10.64898/2026.03.09.710470.