Do historic multiomic profiles improve prediction of postoperative complications?
Biobank metabolomic and proteomic data can predict some long-term disease outcomes, but it is less clear whether historic molecular measurements help predict acute events around surgery. In a new medRxiv preprint, Richard Armstrong and colleagues test whether adding metabolomic and proteomic data improves prediction of postoperative complications after major surgery in UK Biobank.
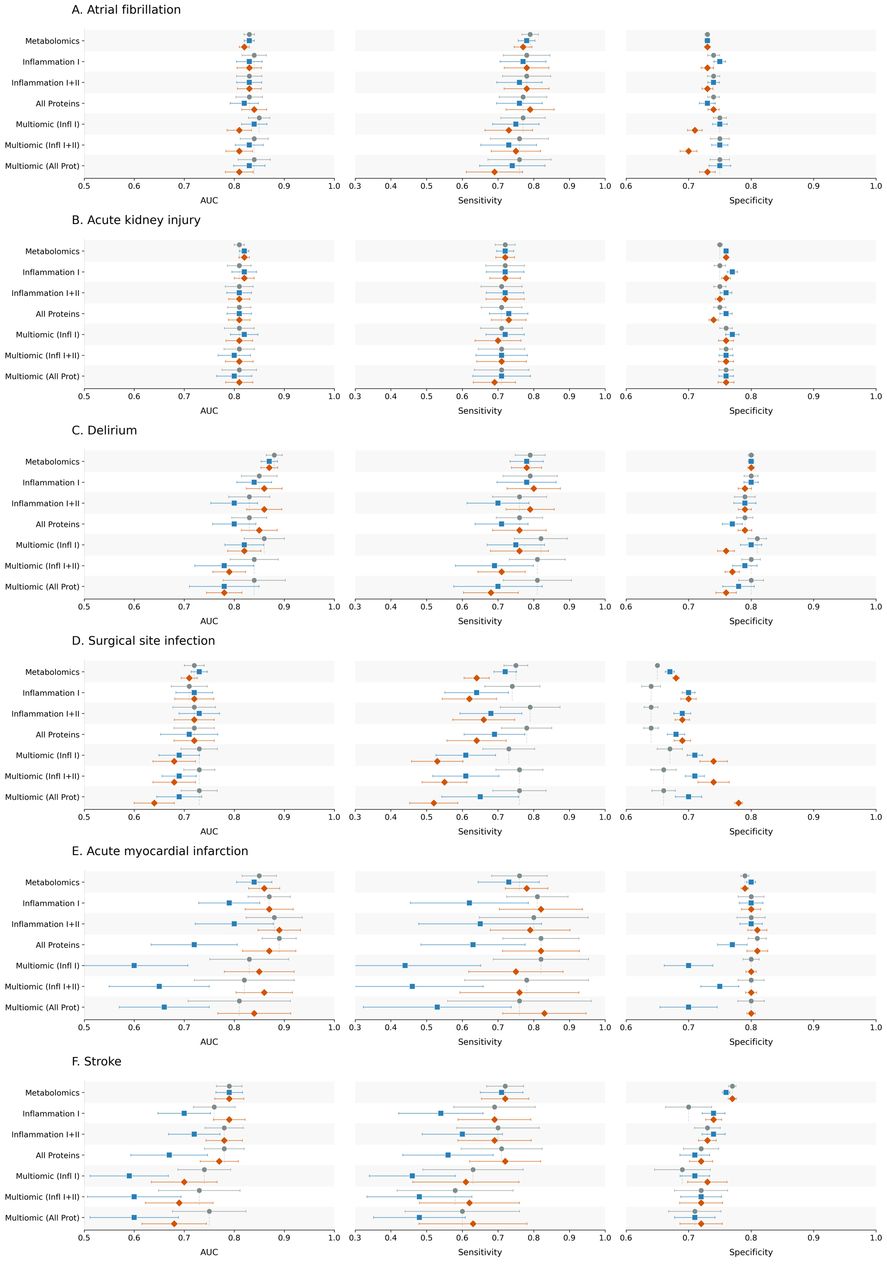
Figure: Model performance across postoperative outcomes and feature sets, comparing clinical baseline models with metabolomic, proteomic and multiomic additions. Source: Armstrong et al., medRxiv, 2026, Fig. 2 (CC BY-NC-ND 4.0).
Background
Preoperative risk prediction usually relies on clinical variables: age, sex, comorbidities, previous diagnoses, procedure type and related information. High-dimensional molecular measurements might capture additional biological vulnerability, but there is a practical question: if those samples were collected months or years before surgery, do they add useful predictive information at the point of care?
This preprint addresses that question for several major postoperative outcomes.
What the study did
The authors analysed 158,156 UK Biobank participants who underwent major surgery. The primary outcomes were postoperative atrial fibrillation, acute kidney injury, acute myocardial infarction, delirium, stroke and surgical site infection.
They compared prediction models based on standard clinical variables with models that added metabolomic data, proteomic data, or both. Because omic subsets were much smaller than the full clinical dataset, the study also used transfer learning from related non-postoperative outcome domains to improve model stability.
What this adds
Baseline clinical models performed reasonably well, with reported Area Under the Receiver Operating Curve (AUROC) values between 0.72 and 0.88. Adding historic metabolomic or proteomic data did not provide a clinically meaningful improvement for the postoperative phenotypes studied.
Transfer learning improved performance and stability in some settings, but still did not outperform the baseline clinical models. The authors interpret this as evidence that long-term biological risk captured by historic omics may already be reflected in routine clinical information by the time someone reaches surgery.
Why this matters
This is a useful negative result. It does not mean omics are irrelevant to surgical outcomes; perioperative or disease-specific molecular measurements may still be valuable. However, it suggests that historic biobank omics, collected at a temporal distance from surgery, may not add enough predictive value to justify routine inclusion in preoperative risk models for these outcomes.
The work is currently a preprint and has not yet been peer reviewed. The small numbers of cases with proteomic or multiomic data for some outcomes are an important limitation, so the conclusion is best read as evidence about this dataset and design rather than a final statement on all perioperative omics.
Preprint
Armstrong RA, Yousefi P, Gibbison B, Khandaker GM, Gaunt TR. Using multiomic data to predict postoperative complications after major surgery in the UK Biobank cohort. medRxiv (posted 2026-03-11). DOI: 10.64898/2026.03.10.26348039.